Define okazaki fragment1/23/2024 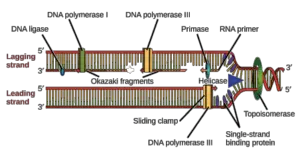
DNA polymerase III is the main DNA replication enzyme in prokaryotes, and it has a much higher affinity for deoxynucleotides than the other two types of DNA polymerase and is therefore able to add nucleotides much more rapidly. DNA polymerase II also plays roles in DNA repair. the removal of the RNA primer and the "stitching up" of Okazaki fragments to form a continuous strand). DNA polymerase I has functions in DNA repair as well as the maturation of Okazaki fragments (i.e. These are very imaginatively named DNA polymerase I, II and III. There are three main types of DNA polymerase in prokaryotes such as E. 
List major DNA polymerases of prokaryotes and There are also single strand DNA-binding proteins that stabilise the unwound strands of DNA, preventing them from "snapping back together." (There's a really bad joke that goes "If I was an enzyme, I'd be helicase so that I can unzip you.") The sliding clamp is a protein that holds DNA Polymerase in place during replication, whereas the clamp loader assembles the clamp onto the DNA using energy from ATP. They cooperate to form what is known as the "replication machine." One of these is helicase, which unzips the double helix. There are several different proteins at the DNA replication fork. Understand the functions of the proteins at the DNA 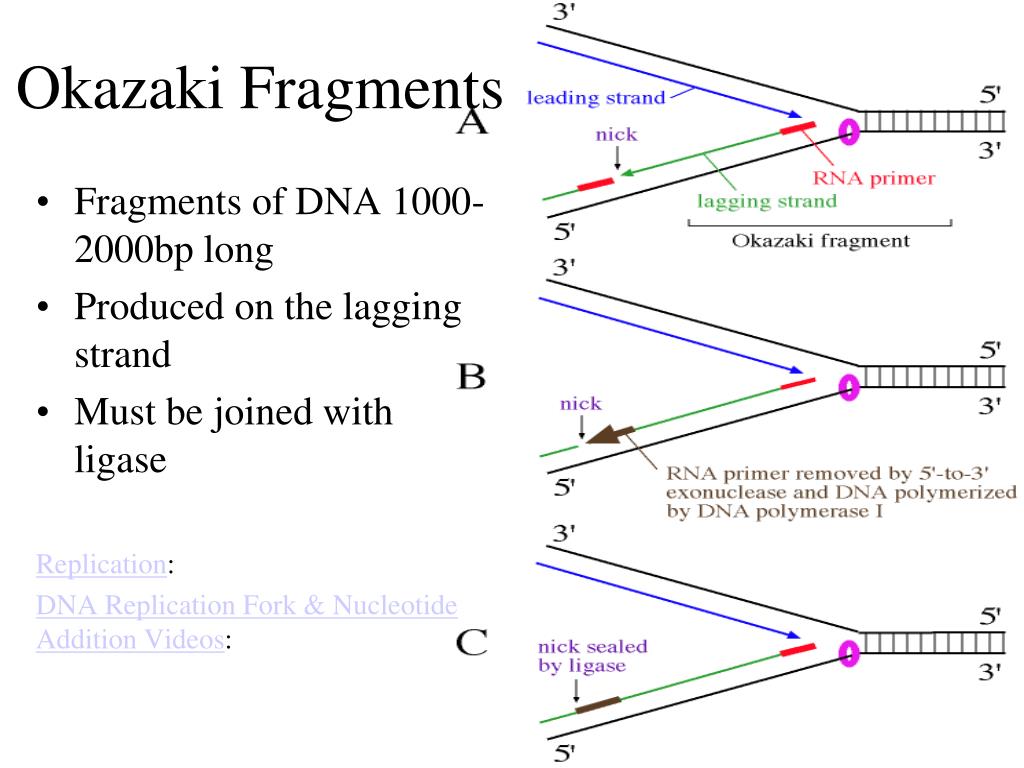
In the case of the lagging strand, most of these RNA primers are removed by RNAse H before DNA polymerase "overwrites" them with DNA. An RNA primer is placed at the start of every leading strand and at the start of every Okazaki fragment on the lagging strand. The cell gets around this by using RNA primers, as RNA doesn't need already existing nucleotides to form a new strand. These fragments are known as Okazaki fragments.Ībout primers- DNA polymerase can only join nucleotides to already existing nucleotides. 
On the other strand, otherwise known as the lagging strand, the DNA is synthesised in little fragments at a time as the helix is opened up. This strand is known as the leading strand. On one strand, this works quite well, and the DNA can be synthesised continuously. Synthesis, however, must run from 5' to 3'. DNA is antiparallel so the two strands are running in opposite directions to each other, and thus synthesis must take place in opposite directions as well. Understand the mechanism of leading and lagging strandĪfter the DNA has been opened up to reveal the replication fork, DNA synthesis takes place on both strands. Okazaki fragment: Fragments of DNA that are formed on the lagging strand. The two DNA strands are separated in this area with DNA replication taking place on both. Replication fork: The area where the DNA is currently being replicated. Eukaryotic chromosomes often have multiple origins.īidirectional: Proceeding in two directions at once. Origin: The place at which DNA replication takes place. This means that it is semiconservative (as opposed to conservative in which the entire daughter molecule would be made of original DNA). Semiconservative: When DNA replicates, each daughter molecule of DNA has one strand from the original DNA and one new strand. Semiconservative, origin, bidirectional, replication Define the terms describing DNA replication:
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
